- Record: found
- Abstract: found
- Article: not found
Chemoproteomic Profiling of Gut Microbiota-Associated Bile Salt Hydrolase Activity
Read this article at
Abstract

The metagenome of the gut microbiome encodes tremendous potential for biosynthesizing and transforming small-molecule metabolites through the activities of enzymes expressed by intestinal bacteria. Accordingly, elucidating this metabolic network is critical for understanding how the gut microbiota contributes to health and disease. Bile acids, which are first biosynthesized in the liver, are modified in the gut by enzymes expressed by commensal bacteria into secondary bile acids, which regulate myriad host processes, including lipid metabolism, glucose metabolism, and immune homeostasis. The gateway reaction of secondary bile acid biosynthesis is mediated by bile salt hydrolases (BSHs), bacterial cysteine hydrolases whose action precedes other bile acid modifications within the gut. To assess how changes in bile acid metabolism mediated by certain intestinal microbiota impact gut physiology and pathobiology, methods are needed to directly examine the activities of BSHs because they are master regulators of intestinal bile acid metabolism. Here, we developed chemoproteomic tools to profile changes in gut microbiome-associated BSH activity. We showed that these probes can label active BSHs in model microorganisms, including relevant gut anaerobes, and in mouse gut microbiomes. Using these tools, we identified altered BSH activities in a murine model of inflammatory bowel disease, in this case, colitis induced by dextran sodium sulfate, leading to changes in bile acid metabolism that could impact host metabolism and immunity. Importantly, our findings reveal that alterations in BSH enzymatic activities within the gut microbiome do not correlate with changes in gene abundance as determined by metagenomic sequencing, highlighting the utility of chemoproteomic approaches for interrogating the metabolic activities of the gut microbiota.
Abstract
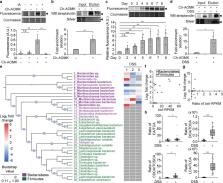
Activity-based profiling of bile salt hydrolase activity using click-chemistry-based chemoproteomics reveals that enzymatic activity increases in a mouse model of colitis.
Related collections
Most cited references17
- Record: found
- Abstract: found
- Article: not found
Immunoglobulin A coating identifies colitogenic bacteria in inflammatory bowel disease.
- Record: found
- Abstract: found
- Article: not found
