- Record: found
- Abstract: found
- Article: found
MACSE v2: Toolkit for the Alignment of Coding Sequences Accounting for Frameshifts and Stop Codons
Read this article at
Abstract
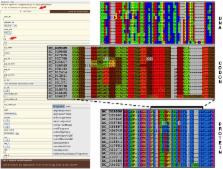
Multiple sequence alignment is a prerequisite for many evolutionary analyses. Multiple Alignment of Coding Sequences (MACSE) is a multiple sequence alignment program that explicitly accounts for the underlying codon structure of protein-coding nucleotide sequences. Its unique characteristic allows building reliable codon alignments even in the presence of frameshifts. This facilitates downstream analyses such as selection pressure estimation based on the ratio of nonsynonymous to synonymous substitutions. Here, we present MACSE v2, a major update with an improved version of the initial algorithm enriched with a complete toolkit to handle multiple alignments of protein-coding sequences. A graphical interface now provides user-friendly access to the different subprograms.
Related collections
Most cited references11


- Record: found
- Abstract: found
- Article: found
Sex-Biased Transcriptome Evolution in Drosophila
- Record: found
- Abstract: found
- Article: not found