- Record: found
- Abstract: found
- Article: found
Interactive Tree of Life (iTOL) v6: recent updates to the phylogenetic tree display and annotation tool

Read this article at
Abstract
The Interactive Tree Of Life ( https://itol.embl.de) is an online tool for the management, display, annotation and manipulation of phylogenetic and other trees. It is freely available and open to everyone. iTOL version 6 introduces a modernized and completely rewritten user interface, together with numerous new features. A new dataset type has been introduced (colored/labeled ranges), greatly upgrading the functionality of the previous simple colored range annotation function. Additional annotation options have been implemented for several existing dataset types. Dataset template files now support simple assignment of annotations to multiple tree nodes through substring matching, including full regular expression support. Node metadata handling has been greatly extended with novel display and exporting options, and it can now be edited interactively or bulk updated through annotation files. Tree labels can be displayed using multiple simultaneous font styles, with precise positioning, sizing and styling of each individual label part. Various bulk label editing functions have been implemented, simplifying large scale changes of all tree node labels. iTOL’s automatic taxonomy assignment functions now support trees based on the Genome Taxonomy Database (GTDB), in addition to the NCBI taxonomy. The functionality of the optional user account pages has been expanded, simplifying the management, navigation and sharing of projects and trees. iTOL currently handles more than one and a half million trees from >130 000 individual user accounts.
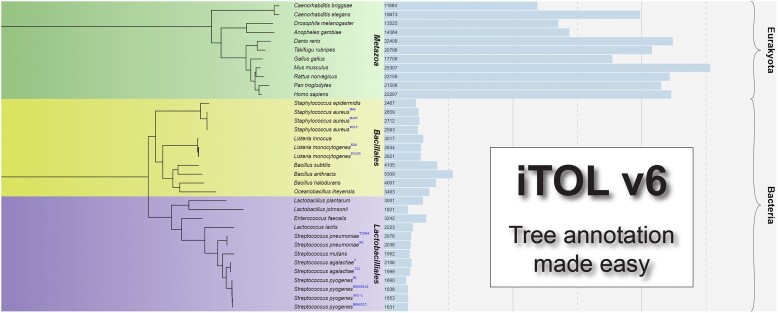
Graphical Abstract
Related collections
Most cited references20

- Record: found
- Abstract: found
- Article: found
Interactive Tree Of Life (iTOL) v5: an online tool for phylogenetic tree display and annotation

- Record: found
- Abstract: found
- Article: found
ETE 3: Reconstruction, Analysis, and Visualization of Phylogenomic Data
- Record: found
- Abstract: found
- Article: not found