- Record: found
- Abstract: found
- Article: found
Integrated Analysis of lncRNA–Mediated ceRNA Network in Lung Adenocarcinoma
Read this article at
Abstract
Background
A growing body of evidence indicates that long non-coding RNAs (lncRNAs) can act as competitive endogenous RNAs (ceRNAs) to bind to microRNAs (miRNAs), thereby affecting and regulating the expression of target genes. The lncRNA–miRNA–mRNA ceRNA network has been theorized to play an indispensable role in many types of tumors. However, the role of the lncRNA-related ceRNA regulatory network in lung adenocarcinoma (LUAD) remains unclear.
Methods
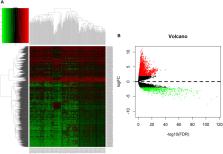
We downloaded the RNAseq and miRNAseq data of LUAD from The Cancer Genome Atlas (TCGA) data portal and identified differentially expressed lncRNAs (DElncRNAs), differentially expressed miRNAs (DEmiRNAs), and differentially expressed mRNAs (DEmRNAs) between LUAD and corresponding paracancerous tissues by using the edgeR package of R software. We constructed the lncRNA–miRNA–mRNA ceRNA network by using Cytoscape (version 3.7.2) on the basis of the interaction generated from the miRcode, miRTarBase, miRDB, and TargetScan databases. Gene Ontology (GO) annotation and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway analysis were performed with DAVID 6.8 bioinformatics resources and plotted by using the ggplot2 package in R. The effect of genes on LUAD prognosis was assessed by applying the survival package in R in accordance with the Kaplan–Meier curve.
Results
In total, 1645 DElncRNAs, 117 DEmiRNAs, and 2729 DEmRNAs were identified in LUAD. The LUAD-specific ceRNA network was composed of 157 nodes and 378 edges (329 DElncRNA–DEmiRNA interactions and 49 DEmiRNA–DEmRNA interactions). GO and KEGG pathway annotations suggested that the LUAD-specific ceRNA network was related to tumor-related molecular functions and pathways. Seven lncRNAs (DISC1-IT1, SYNPR-AS1, H19, LINC00460, LINC00518, DSCR10, and STEAP2-AS1), one miRNA (hsa-mir-31), and 16 mRNAs (ATAD2, OSCAR, KIF23, E2F7, PFKP, MCM4, CEP55, CBX2, CCNE1, CLSPN, CCNB1, CDC25A, EZH2, CHEK1, SLC7A11, and PBK) were revealed to be significantly correlated with overall survival.
Related collections
Most cited references29
- Record: found
- Abstract: found
- Article: not found
Microarray analysis shows that some microRNAs downregulate large numbers of target mRNAs.
- Record: found
- Abstract: found
- Article: not found
Switching from repression to activation: microRNAs can up-regulate translation.
- Record: found
- Abstract: found
- Article: found