- Record: found
- Abstract: found
- Article: found
Kaptive Web: User-Friendly Capsule and Lipopolysaccharide Serotype Prediction for Klebsiella Genomes
Read this article at
ABSTRACT
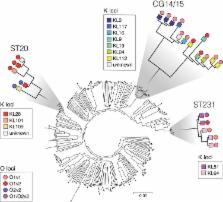
As whole-genome sequencing becomes an established component of the microbiologist's toolbox, it is imperative that researchers, clinical microbiologists, and public health professionals have access to genomic analysis tools for the rapid extraction of epidemiologically and clinically relevant information. For the Gram-negative hospital pathogens such as Klebsiella pneumoniae, initial efforts have focused on the detection and surveillance of antimicrobial resistance genes and clones. However, with the resurgence of interest in alternative infection control strategies targeting Klebsiella surface polysaccharides, the ability to extract information about these antigens is increasingly important. Here we present Kaptive Web, an online tool for the rapid typing of Klebsiella K and O loci, which encode the polysaccharide capsule and lipopolysaccharide O antigen, respectively. Kaptive Web enables users to upload and analyze genome assemblies in a web browser. The results can be downloaded in tabular format or explored in detail via the graphical interface, making it accessible for users at all levels of computational expertise. We demonstrate Kaptive Web's utility by analyzing >500 K. pneumoniae genomes. We identify extensive K and O locus diversity among 201 genomes belonging to the carbapenemase-associated clonal group 258 (25 K and 6 O loci). The characterization of a further 309 genomes indicated that such diversity is common among the multidrug-resistant clones and that these loci represent useful epidemiological markers for strain subtyping. These findings reinforce the need for rapid, reliable, and accessible typing methods such as Kaptive Web. Kaptive Web is available for use at http://kaptive.holtlab.net/, and the source code is available at https://github.com/kelwyres/Kaptive-Web.
Related collections
Most cited references49
- Record: found
- Abstract: found
- Article: found
Completing bacterial genome assemblies with multiplex MinION sequencing

- Record: found
- Abstract: found
- Article: found
Identification of Klebsiella capsule synthesis loci from whole genome data
- Record: found
- Abstract: found
- Article: not found