- Record: found
- Abstract: not found
- Article: not found
Computational methods in glaucoma research: Current status and future outlook
December 2023
December 2023
Read this article at
ScienceOpenPublisher
There is no author summary for this article yet. Authors can add summaries to their articles on ScienceOpen to make them more accessible to a non-specialist audience.
Related collections
Most cited references185

- Record: found
- Abstract: found
- Article: found
AlphaFold Protein Structure Database: massively expanding the structural coverage of protein-sequence space with high-accuracy models
Mihaly Varadi, Stephen Anyango, Mandar Deshpande … (2021)

- Record: found
- Abstract: found
- Article: found
The Ensembl Variant Effect Predictor
William McLaren, Laurent Gil, Sarah Hunt … (2016)
- Record: found
- Abstract: found
- Article: found
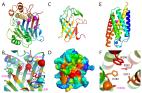
Accurate prediction of protein structures and interactions using a 3-track neural network
Minkyung Baek, Frank DiMaio, Ivan V Anishchenko … (2022)
Author and article information
Comments
Comment on this article
scite_
0
0
0
0
 Smart Citations
Smart Citations0
0
0
0
Citing PublicationsSupportingMentioningContrasting
See how this article has been cited at scite.ai
scite shows how a scientific paper has been cited by providing the context of the citation, a classification describing whether it supports, mentions, or contrasts the cited claim, and a label indicating in which section the citation was made.
