- Record: found
- Abstract: found
- Article: found
The role of the Mediator complex in fungal pathogenesis and response to antifungal agents
Read this article at
Abstract
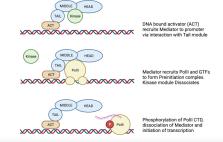
Mediator is a complex of polypeptides that plays a central role in the recruitment of RNA polymerase II to promoters and subsequent transcriptional activation in eukaryotic organisms. Studies have now shown that Mediator has a role in regulating expression of genes implicated in virulence and antifungal drug resistance in pathogenic fungi. The roles of specific Mediator subunits have been investigated in several species of pathogenic fungi, particularly in the most pathogenic yeast Candida albicans. Uniquely, pathogenic yeast also present several interesting examples of divergence in Mediator structure and function, most notably in C. glabrata, which possesses two orthologues of Med15, and in C. albicans, which has a massively expanded family of Med2 orthologues known as the TLO gene family. This review highlights specific examples of recent progress in characterizing the role of Mediator in pathogenic fungi.
Related collections
Most cited references63

- Record: found
- Abstract: found
- Article: found
Highly accurate protein structure prediction with AlphaFold

- Record: found
- Abstract: found
- Article: found
AlphaFold Protein Structure Database: massively expanding the structural coverage of protein-sequence space with high-accuracy models

- Record: found
- Abstract: found
- Article: found