- Record: found
- Abstract: found
- Article: found
AI-Driven Multiscale Simulations Illuminate Mechanisms of SARS-CoV-2 Spike Dynamics
Read this article at
Abstract
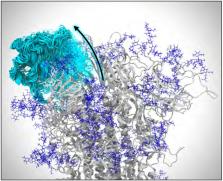
We develop a generalizable AI-driven workflow that leverages heterogeneous HPC resources to explore the time-dependent dynamics of molecular systems. We use this workflow to investigate the mechanisms of infectivity of the SARS-CoV-2 spike protein, the main viral infection machinery. Our workflow enables more efficient investigation of spike dynamics in a variety of complex environments, including within a complete SARS-CoV-2 viral envelope simulation, which contains 305 million atoms and shows strong scaling on ORNL Summit using NAMD. We present several novel scientific discoveries, including the elucidation of the spike’s full glycan shield, the role of spike glycans in modulating the infectivity of the virus, and the characterization of the flexible interactions between the spike and the human ACE2 receptor. We also demonstrate how AI can accelerate conformational sampling across different systems and pave the way for the future application of such methods to additional studies in SARS-CoV-2 and other molecular systems.
Related collections
Most cited references77

- Record: found
- Abstract: found
- Article: found
Cryo-EM structure of the 2019-nCoV spike in the prefusion conformation
- Record: found
- Abstract: found
- Article: not found
Structure, Function, and Antigenicity of the SARS-CoV-2 Spike Glycoprotein
- Record: found
- Abstract: not found
- Article: not found