- Record: found
- Abstract: found
- Article: found
YUCCA6 over-expression demonstrates auxin function in delaying leaf senescence in Arabidopsis thaliana
Read this article at
Abstract
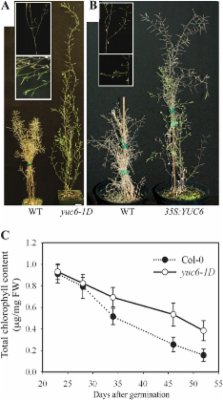
The Arabidopsis thaliana YUCCA family of flavin monooxygenase proteins catalyses a rate-limiting step in de novo auxin biosynthesis. A YUCCA6 activation mutant, yuc6-1D, has been shown to contain an elevated free IAA level and to display typical high-auxin phenotypes. It is reported here that Arabidopsis plants over-expressing YUCCA6, such as the yuc6-1D activation mutant and 35S: YUC6 transgenic plants, displayed dramatic longevity. In addition, plants over-expressing YUCCA6 exhibited classical, delayed dark-induced and hormone-induced senescence in assays using detached rosette leaves. However, plants over-expressing an allele of YUCCA6, that carries mutations in the NADPH cofactor binding site, exhibited neither delayed leaf senescence phenotypes nor phenotypes typical of auxin overproduction. When the level of free IAA was reduced in yuc6-1D by conjugation to lysine, yuc6-1D leaves senesced at a rate similar to the wild-type leaves. Dark-induced senescence in detached leaves was accompanied by a decrease in their free IAA content, by the reduced expression of auxin biosynthesis enzymes such as YUCCA1 and YUCCA6 that increase cellular free IAA levels, and by the increased expression of auxin-conjugating enzymes encoded by the GH3 genes that reduce the cellular free auxin levels. Reduced transcript abundances of SAG12, NAC1, and NAC6 during senescence in yuc6-1D compared with the wild type suggested that auxin delays senescence by directly or indirectly regulating the expression of senescence-associated genes.
Related collections
Most cited references57
- Record: found
- Abstract: found
- Article: not found
TAA1-mediated auxin biosynthesis is essential for hormone crosstalk and plant development.
- Record: found
- Abstract: found
- Article: not found
MicroRNA directs mRNA cleavage of the transcription factor NAC1 to downregulate auxin signals for arabidopsis lateral root development.
- Record: found
- Abstract: found
- Article: not found