- Record: found
- Abstract: found
- Article: found
Detection of 224 candidate structured RNAs by comparative analysis of specific subsets of intergenic regions
Read this article at
Abstract
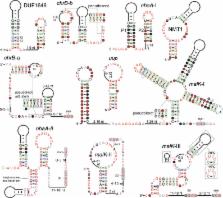
The discovery of structured non-coding RNAs (ncRNAs) in bacteria can reveal new facets of biology and biochemistry. Comparative genomics analyses executed by powerful computer algorithms have successfully been used to uncover many novel bacterial ncRNA classes in recent years. However, this general search strategy favors the discovery of more common ncRNA classes, whereas progressively rarer classes are correspondingly more difficult to identify. In the current study, we confront this problem by devising several methods to select subsets of intergenic regions that can concentrate these rare RNA classes, thereby increasing the probability that comparative sequence analysis approaches will reveal their existence. By implementing these methods, we discovered 224 novel ncRNA classes, which include ROOL RNA, an RNA class averaging 581 nt and present in multiple phyla, several highly conserved and widespread ncRNA classes with properties that suggest sophisticated biochemical functions and a multitude of putative cis-regulatory RNA classes involved in a variety of biological processes. We expect that further research on these newly found RNA classes will reveal additional aspects of novel biology, and allow for greater insights into the biochemistry performed by ncRNAs.
Related collections
Most cited references51
- Record: found
- Abstract: found
- Article: not found
LIBSVM: A library for support vector machines
- Record: found
- Abstract: found
- Article: not found
Fast and reliable prediction of noncoding RNAs.
- Record: found
- Abstract: found
- Article: not found