- Record: found
- Abstract: found
- Article: found
First Detection of Bat Astroviruses (BtAstVs) among Bats in Poland: The Genetic BtAstVs Diversity Reveals Multiple Co-Infection of Bats with Different Strains
Read this article at
Abstract
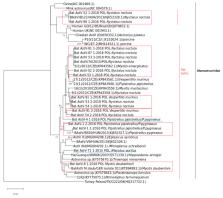
Background: Astroviruses (AstVs) are common pathogens of a wide range of animal hosts, including mammals and avians, causing gastrointestinal diseases, mainly gastroenteritis and diarrhea. They prompt a significant health problem in newborns and young children and economic losses in the poultry sector and mink farms. Recent studies revealed a growing number of bat species carrying astroviruses with a noticeable prevalence and diversity. Here, we demonstrate the first detection of bat astroviruses (BtAstVs) circulating in the population of insectivorous bats in the territory of Poland. Results: Genetically diverse BtAstVs (n = 18) were found with a varying degree of bat species specificity in five out of 15 bat species in Poland previously recognized as BtAstV hosts. Astroviral RNA was found in 12 out of 98 (12.2%, 95% CI 7.1–20.2) bat intestines, six bat kidneys (6.1%, 95% CI 2.8–12.7) and two bat livers (2.0%, 95% CI 0.4–7.1). Deep sequencing of the astroviral RNA-dependent RNA polymerase (RdRp) region revealed co-infections in five single bat individuals with highly distinct astrovirus strains. Conclusions: The detection of highly distinct bat astroviruses in Polish bats favors virus recombination and the generation of novel divergent AstVs and creates a potential risk of virus transmission to domestic animals and humans in the country. These findings provide a new insight into molecular epidemiology, prevalence of astroviruses in European bat populations and the risk of interspecies transmission to other animals including humans.
Related collections
Most cited references39
- Record: found
- Abstract: found
- Article: not found
MEGA4: Molecular Evolutionary Genetics Analysis (MEGA) software version 4.0.
- Record: found
- Abstract: found
- Article: not found
Bats: important reservoir hosts of emerging viruses.
- Record: found
- Abstract: found
- Article: not found