- Record: found
- Abstract: found
- Article: found
Protocol to decode the role of transcriptionally active microbes in SARS-CoV-2-positive patients using an RNA-seq-based approach

Read this article at
Summary
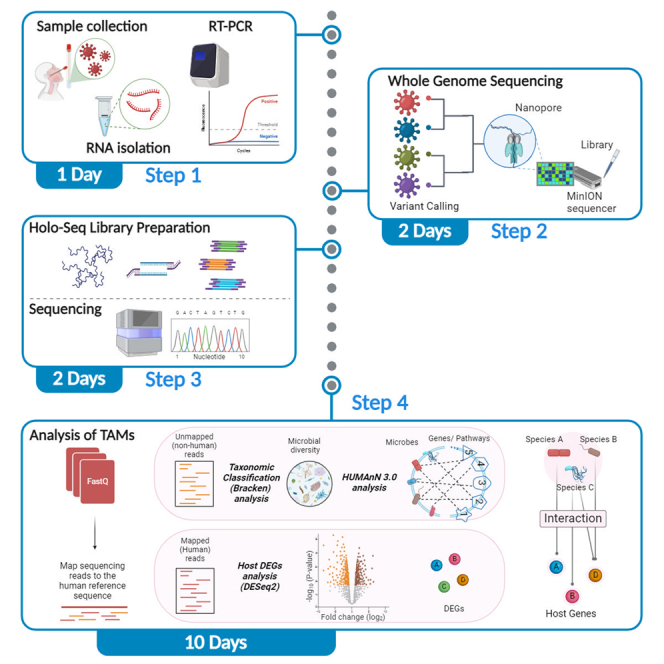
The elucidation of the role of microorganisms in human infections has been hindered by difficulties using conventional culture-based techniques. Here, we present a protocol for the investigation of transcriptionally active microbes (TAMs) using an RNA sequencing (RNA-seq)-based approach. We describe the steps for RNA isolation, viral genome sequencing, RNA-seq library preparation, and metatranscriptomic and transcriptomic analysis. This protocol permits a comprehensive evaluation of TAMs’ contributions to the differential severity of infectious diseases, with a particular focus on diseases such as COVID-19.
For complete details on the use and execution of this protocol, please refer to Devi et al. 1
Graphical abstract
Highlights
-
•
A protocol to characterize transcriptionally active microbes (TAMs) using RNA-seq
-
•
Steps for RNA-seq library preparation, sequencing, and metatranscriptomic analysis
-
•
Detailed procedure for taxonomic and functional classification of microbes
-
•
Correlation between bacterial species and the expressed host genes
Abstract
Publisher’s note: Undertaking any experimental protocol requires adherence to local institutional guidelines for laboratory safety and ethics.
Abstract
The elucidation of the role of microorganisms in human infections has been hindered by difficulties using conventional culture-based techniques. Here, we present a protocol for the investigation of transcriptionally active microbes (TAMs) using an RNA sequencing (RNA-seq)-based approach. We describe the steps for RNA isolation, viral genome sequencing, RNA-seq library preparation, and metatranscriptomic and transcriptomic analysis. This protocol permits a comprehensive evaluation of TAMs’ contributions to the differential severity of infectious diseases, with a particular focus on diseases such as COVID-19.
Related collections
Most cited references14

- Record: found
- Abstract: found
- Article: found
Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2

- Record: found
- Abstract: found
- Article: found
Trimmomatic: a flexible trimmer for Illumina sequence data
- Record: found
- Abstract: found
- Article: not found