- Record: found
- Abstract: found
- Article: found
Exploring the Pharmacological Mechanism of Duhuo Jisheng Decoction in Treating Osteoporosis Based on Network Pharmacology
Read this article at
Abstract
Objective
The purpose of this work is to study the mechanism of action of Duhuo Jisheng Decoction (DHJSD) in the treatment of osteoporosis based on the methods of bioinformatics and network pharmacology.
Methods
In this study, the active compounds of each medicinal ingredient of DHJSD and their corresponding targets were obtained from TCMSP database. Osteoporosis was treated as search query in GeneCards, MalaCards, DisGeNET, Therapeutic Target Database (TTD), Comparative Toxicogenomics Database (CTD), and OMIM databases to obtain disease-related genes. The overlapping targets of DHJSD and osteoporosis were identified, and then GO and KEGG enrichment analysis were performed. Cytoscape was employed to construct DHJSD-compounds-target genes-osteoporosis network and protein-protein interaction (PPI) network. CytoHubba was utilized to select the hub genes. The activities of binding of hub genes and key components were confirmed by molecular docking.
Results
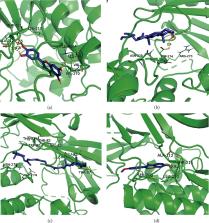
174 active compounds and their 205 related potential targets were identified in DHJSD for the treatment of osteoporosis, including 10 hub genes (AKT1, ALB, IL6, MAPK3, VEGFA, JUN, CASP3, EGFR, MYC, and EGF). Pathway enrichment analysis of target proteins indicated that osteoclast differentiation, AGE-RAGE signaling pathway in diabetic complications, Wnt signaling pathway, MAPK signaling pathway, PI3K-Akt signaling pathway, JAK-STAT signaling pathway, calcium signaling pathway, and TNF signaling pathway were the specifically major pathways regulated by DHJSD against osteoporosis. Further verification based on molecular docking results showed that the small molecule compounds (Quercetin, Kaempferol, Beta-sitosterol, Beta-carotene, and Formononetin) contained in DHJSD generally have excellent binding affinity to the macromolecular target proteins encoded by the top 10 genes.
Related collections
Most cited references68
- Record: found
- Abstract: found
- Article: not found
clusterProfiler: an R package for comparing biological themes among gene clusters.
- Record: found
- Abstract: found
- Article: not found
Cytoscape: a software environment for integrated models of biomolecular interaction networks.
- Record: found
- Abstract: found
- Article: not found