- Record: found
- Abstract: found
- Article: found
The Changing Pattern of Population Structure of Staphylococcus aureus from Bacteremia in China from 2013 to 2016: ST239-030-MRSA Replaced by ST59-t437
Read this article at
Abstract
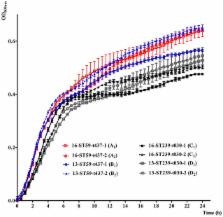
To investigate the epidemiology and genetic structure of Staphylococcus aureus bacteremia in China, a total of 416 isolates from 22 teaching hospitals in 12 cities from 2013 and 2016 were characterized by antibiogram analysis, multilocus sequence typing (MLST), spa typing and staphylococcal cassette chromosome mec (SCC mec) typing. The predominant meticillin-susceptible (MSSA) genotypes in 2013 were ST188 (19.1%), ST7 (8.7%), and ST398 (7.8%), respectively, and they continued to be the main genotypes in 2016. The prevalence of meticillin-resistant S. aureus (MRSA) were 36.5% (66/181) and 36.6% (86/235) in 2013 and 2016, respectively. Interestingly, the susceptibility rates of MRSA to rifampicin and fluoroquinolones increased significantly from 2013 to 2016 ( P < 0.01), and this was associated with changes in genetic structure. ST239-t030-MRSA, the predominant genotype among all MRSAs in 2013 (34.8%), was replaced by ST59-t437-MRSA (15.1%) in 2016. Further analysis revealed that the ST239-t030-MRSA were more resistant to rifampicin, tetracycline and fluoroquinolones than ST59-t437-MRSA ( P < 0.01). To further gain insight into the mechanisms underlying the changes of genetic structure, in vitro competition and fitness measurements were performed. Importantly, ST239-t030-MRSA displayed lower growth rate and lower competitive advantage compared to ST59-t437-MRSA. Together, our findings reveal that fitness advantage of ST59-t437-MRSA over ST239-t030-MRSA may lead to changes in genetic structure and increased susceptibility of MRSA to rifampicin and fluoroquinolones in Chinese patients with S. aureus bacteremia. Our study supports temporal dynamics in MRSA clone diversities, further providing critical insights into the importance of continued monitoring of MRSA.
Related collections
Most cited references25

- Record: found
- Abstract: found
- Article: found
Trends in antimicrobial resistance in bloodstream infection isolates at a large urban hospital in Malawi (1998–2016): a surveillance study
- Record: found
- Abstract: found
- Article: not found
Infectious disease consultation for Staphylococcus aureus bacteremia - A systematic review and meta-analysis.
- Record: found
- Abstract: found
- Article: not found