- Record: found
- Abstract: found
- Article: not found
START lipid/sterol-binding domains are amplified in plants and are predominantly associated with homeodomain transcription factors
Read this article at
Abstract
A survey of proteins containing lipid/sterol-binding StAR-related lipid transfer (START) domains shows that they are amplified in plants and are primarily found within homeodomain (HD) transcription factors.
Abstract
Background
In animals, steroid hormones regulate gene expression by binding to nuclear receptors. Plants lack genes for nuclear receptors, yet genetic evidence from Arabidopsis suggests developmental roles for lipids/sterols analogous to those in animals. In contrast to nuclear receptors, the lipid/sterol-binding StAR-related lipid transfer (START) protein domains are conserved, making them candidates for involvement in both animal and plant lipid/sterol signal transduction.
Results
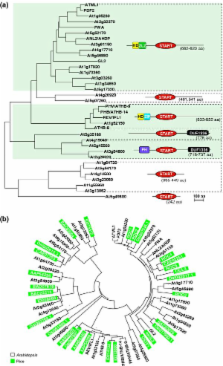
We surveyed putative START domains from the genomes of Arabidopsis, rice, animals, protists and bacteria. START domains are more common in plants than in animals and in plants are primarily found within homeodomain (HD) transcription factors. The largest subfamily of HD-START proteins is characterized by an HD amino-terminal to a plant-specific leucine zipper with an internal loop, whereas in a smaller subfamily the HD precedes a classic leucine zipper. The START domains in plant HD-START proteins are not closely related to those of animals, implying collateral evolution to accommodate organism-specific lipids/sterols. Using crystal structures of mammalian START proteins, we show structural conservation of the mammalian phosphatidylcholine transfer protein (PCTP) START domain in plants, consistent with a common role in lipid transport and metabolism. We also describe putative START-domain proteins from bacteria and unicellular protists.
Conclusions
The majority of START domains in plants belong to a novel class of putative lipid/sterol-binding transcription factors, the HD-START family, which is conserved across the plant kingdom. HD-START proteins are confined to plants, suggesting a mechanism by which lipid/sterol ligands can directly modulate transcription in plants.
Related collections
Most cited references47
- Record: found
- Abstract: found
- Article: not found
The Pfam protein families database.
- Record: found
- Abstract: found
- Article: not found
A comprehensive set of sequence analysis programs for the VAX.
- Record: found
- Abstract: found
- Article: not found
