- Record: found
- Abstract: found
- Article: found
Respiratory viruses interacting with cells: the importance of electrostatics
Read this article at
Abstract
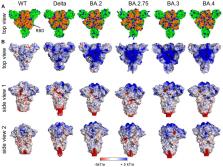
The COVID-19 pandemic has rekindled interest in the molecular mechanisms involved in the early steps of infection of cells by viruses. Compared to SARS-CoV-1 which only caused a relatively small albeit deadly outbreak, SARS-CoV-2 has led to fulminant spread and a full-scale pandemic characterized by efficient virus transmission worldwide within a very short time. Moreover, the mutations the virus acquired over the many months of virus transmission, particularly those seen in the Omicron variant, have turned out to result in an even more transmissible virus. Here, we focus on the early events of virus infection of cells. We review evidence that the first decisive step in this process is the electrostatic interaction of the spike protein with heparan sulfate chains present on the surface of target cells: Patches of cationic amino acids located on the surface of the spike protein can interact intimately with the negatively charged heparan sulfate chains, which results in the binding of the virion to the cell surface. In a second step, the specific interaction of the receptor binding domain (RBD) within the spike with the angiotensin-converting enzyme 2 (ACE2) receptor leads to the uptake of bound virions into the cell. We show that these events can be expressed as a semi-quantitative model by calculating the surface potential of different spike proteins using the Adaptive Poison-Boltzmann-Solver (APBS). This software allows visualization of the positive surface potential caused by the cationic patches, which increased markedly from the original Wuhan strain of SARS-CoV-2 to the Omicron variant. The surface potential thus enhanced leads to a much stronger binding of the Omicron variant as compared to the original wild-type virus. At the same time, data taken from the literature demonstrate that the interaction of the RBD of the spike protein with the ACE2 receptor remains constant within the limits of error. Finally, we briefly digress to other viruses and show the usefulness of these electrostatic processes and calculations for cell-virus interactions more generally.
Related collections
Most cited references123
- Record: found
- Abstract: found
- Article: not found
SARS-CoV-2 Cell Entry Depends on ACE2 and TMPRSS2 and Is Blocked by a Clinically Proven Protease Inhibitor

- Record: found
- Abstract: found
- Article: found
Cryo-EM structure of the 2019-nCoV spike in the prefusion conformation
- Record: found
- Abstract: found
- Article: not found