- Record: found
- Abstract: found
- Article: found
GWIPS-viz: 2018 update
Read this article at
Abstract
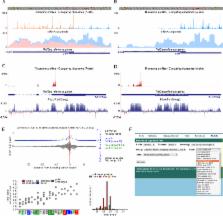
The GWIPS-viz browser ( http://gwips.ucc.ie/) is an on-line genome browser which is tailored for exploring ribosome profiling (Ribo-seq) data. Since its publication in 2014, GWIPS-viz provides Ribo-seq data for an additional 14 genomes bringing the current total to 23. The integration of new Ribo-seq data has been automated thereby increasing the number of available tracks to 1792, a 10-fold increase in the last three years. The increase is particularly substantial for data derived from human sources. Following user requests, we added the functionality to download these tracks in bigWig format. We also incorporated new types of data (e.g. TCP-seq) as well as auxiliary tracks from other sources that help with the interpretation of Ribo-seq data. Improvements in the visualization of the data have been carried out particularly for bacterial genomes where the Ribo-seq data are now shown in a strand specific manner. For higher eukaryotic datasets, we provide characteristics of individual datasets using the RUST program which includes the triplet periodicity, sequencing biases and relative inferred A-site dwell times. This information can be used for assessing the quality of Ribo-seq datasets. To improve the power of the signal, we aggregate Ribo-seq data from several studies into Global aggregate tracks for each genome.
Related collections
Most cited references111

- Record: found
- Abstract: found
- Article: found
The UCSC Genome Browser database: update 2011
- Record: found
- Abstract: found
- Article: not found
Poly(A)-tail profiling reveals an embryonic switch in translational control
- Record: found
- Abstract: found
- Article: not found