- Record: found
- Abstract: found
- Article: found
circAtlas 3.0: a gateway to 3 million curated vertebrate circular RNAs based on a standardized nomenclature scheme

Read this article at
Abstract
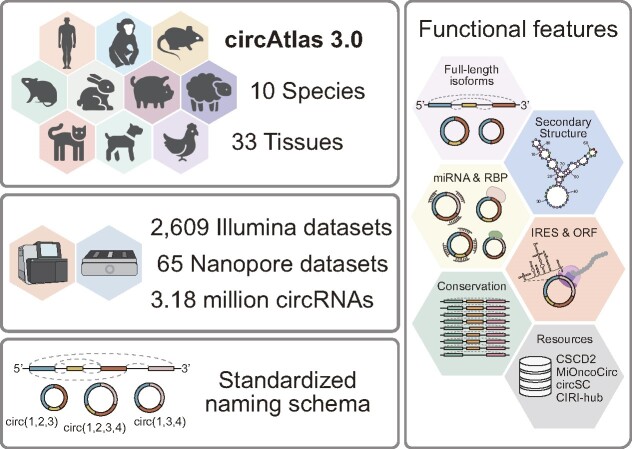
Recent studies have demonstrated the important regulatory role of circRNAs, but an in-depth understanding of the comprehensive landscape of circRNAs across various species still remains unexplored. The current circRNA databases are often species-restricted or based on outdated datasets. To address this challenge, we have developed the circAtlas 3.0 database, which contains a rich collection of 2674 circRNA sequencing datasets, curated to delineate the landscape of circRNAs within 33 distinct tissues spanning 10 vertebrate species. Notably, circAtlas 3.0 represents a substantial advancement over its precursor, circAtlas 2.0, with the number of cataloged circRNAs escalating from 1 007 087 to 3 179 560, with 2 527 528 of them being reconstructed into full-length isoforms. circAtlas 3.0 also introduces several notable enhancements, including: (i) integration of both Illumina and Nanopore sequencing datasets to detect circRNAs of extended lengths; (ii) employment of a standardized nomenclature scheme for circRNAs, providing information of the host gene and full-length circular exons; (iii) inclusion of clinical cancer samples to explore the biological function of circRNAs within the context of cancer and (iv) links to other useful resources to enable user-friendly analysis of target circRNAs. The updated circAtlas 3.0 provides an important platform for exploring the evolution and biological implications of vertebrate circRNAs, and is freely available at http://circatlas.biols.ac.cn and https://ngdc.cncb.ac.cn/circatlas.
Graphical Abstract
Related collections
Most cited references63
- Record: found
- Abstract: found
- Article: not found
Circular RNAs are a large class of animal RNAs with regulatory potency.
- Record: found
- Abstract: found
- Article: not found
Natural RNA circles function as efficient microRNA sponges.

- Record: found
- Abstract: found
- Article: found