- Record: found
- Abstract: found
- Article: found
Therapeutic target database update 2018: enriched resource for facilitating bench-to-clinic research of targeted therapeutics
Read this article at
Abstract
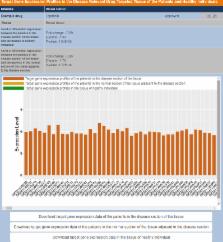
Extensive efforts have been directed at the discovery, investigation and clinical monitoring of targeted therapeutics. These efforts may be facilitated by the convenient access of the genetic, proteomic, interactive and other aspects of the therapeutic targets. Here, we describe an update of the Therapeutic target database (TTD) previously featured in NAR. This update includes: (i) 2000 drug resistance mutations in 83 targets and 104 target/drug regulatory genes, which are resistant to 228 drugs targeting 63 diseases (49 targets of 61 drugs with patient prevalence data); (ii) differential expression profiles of 758 targets in the disease-relevant drug-targeted tissue of 12 615 patients of 70 diseases; (iii) expression profiles of 629 targets in the non-targeted tissues of 2565 healthy individuals; (iv) 1008 target combinations of 1764 drugs and the 1604 target combination of 664 multi-target drugs; (v) additional 48 successful, 398 clinical trial and 21 research targets, 473 approved, 812 clinical trial and 1120 experimental drugs, and (vi) ICD-10-CM and ICD-9-CM codes for additional 482 targets and 262 drugs against 98 disease conditions. This update makes TTD more useful for facilitating the patient focused research, discovery and clinical investigations of the targeted therapeutics. TTD is accessible at http://bidd.nus.edu.sg/group/ttd/ttd.asp.
Related collections
Most cited references62
- Record: found
- Abstract: found
- Article: not found
Basic local alignment search tool.

- Record: found
- Abstract: found
- Article: found
NCBI GEO: archive for functional genomics data sets—update

- Record: found
- Abstract: found
- Article: found