- Record: found
- Abstract: found
- Article: found
Microcollinearity between autopolyploid sugarcane and diploid sorghum genomes
Read this article at
Abstract
Background
Sugarcane ( Saccharum spp.) has become an increasingly important crop for its leading role in biofuel production. The high sugar content species S. officinarum is an octoploid without known diploid or tetraploid progenitors. Commercial sugarcane cultivars are hybrids between S. officinarum and wild species S. spontaneum with ploidy at ~12×. The complex autopolyploid sugarcane genome has not been characterized at the DNA sequence level.
Results
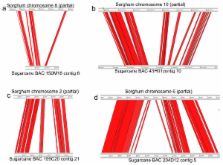
The microsynteny between sugarcane and sorghum was assessed by comparing 454 pyrosequences of 20 sugarcane bacterial artificial chromosomes (BACs) with sorghum sequences. These 20 BACs were selected by hybridization of 1961 single copy sorghum overgo probes to the sugarcane BAC library with one sugarcane BAC corresponding to each of the 20 sorghum chromosome arms. The genic regions of the sugarcane BACs shared an average of 95.2% sequence identity with sorghum, and the sorghum genome was used as a template to order sequence contigs covering 78.2% of the 20 BAC sequences. About 53.1% of the sugarcane BAC sequences are aligned with sorghum sequence. The unaligned regions contain non-coding and repetitive sequences. Within the aligned sequences, 209 genes were annotated in sugarcane and 202 in sorghum. Seventeen genes appeared to be sugarcane-specific and all validated by sugarcane ESTs, while 12 appeared sorghum-specific but only one validated by sorghum ESTs. Twelve of the 17 sugarcane-specific genes have no match in the non-redundant protein database in GenBank, perhaps encoding proteins for sugarcane-specific processes. The sorghum orthologous regions appeared to have expanded relative to sugarcane, mostly by the increase of retrotransposons.
Conclusions
The sugarcane and sorghum genomes are mostly collinear in the genic regions, and the sorghum genome can be used as a template for assembling much of the genic DNA of the autopolyploid sugarcane genome. The comparable gene density between sugarcane BACs and corresponding sorghum sequences defied the notion that polyploidy species might have faster pace of gene loss due to the redundancy of multiple alleles at each locus.
Related collections
Most cited references27
- Record: found
- Abstract: found
- Article: not found
The Sorghum bicolor genome and the diversification of grasses.
- Record: found
- Abstract: found
- Article: not found
Ecological intensification of cereal production systems: yield potential, soil quality, and precision agriculture.
- Record: found
- Abstract: found
- Article: not found