- Record: found
- Abstract: found
- Article: found
Genetic alteration and clinical significance of SUMOylation regulators in multiple cancer types
Read this article at
Abstract
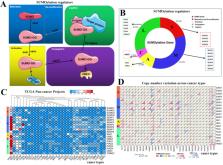
The purpose of this study was to investigate the genetic variation, gene expression differences, and clinical significance of SUMOylation regulators in pan-cancers. Based on previous studies, we gained a better understanding of the biological process of SUMOylation and the status of current research. In the present study, we employed a wide range of bioinformatics methods. We used genetic variation and mRNA expression data in the Cancer Genome Atlas (TCGA) to construct a panoramic view of the single nucleotide variants, copy number variants, and gene expression changes in SUMOylation regulators in various tumors. Subsequently, we used the String website and the Cytoscape tool to construct the PPI network between these regulators. We used the GSCALite website to determine the relationship between these regulators and cancer pathways and drug sensitivity. We constructed images of co-expression between these regulators using the R programming language. Using clinical data from TCGA, we performed hazard ratio analysis for these regulators in pan-cancer. Most importantly, we used these regulators to successfully establish risk signatures related to patient prognosis in multiple tumors. Finally, in KIRC, we conducted gene-set enrichment analysis (GSEA) of the five molecules in its risk signatures. We found that these five molecules are involved in multiple cancer pathways. In short, we have comprehensively interpreted the detailed biological process of SUMOylation at the genetic level for the first time, successfully constructed multiple risk signatures, and conducted GSEA in KIRC. We believe that these findings provide credible and valuable information that is relevant for future clinical diagnoses and scientific research.
Related collections
Most cited references44
- Record: found
- Abstract: found
- Article: not found
Cytoscape: a software environment for integrated models of biomolecular interaction networks.
- Record: found
- Abstract: found
- Article: not found
Cancer statistics, 2019

- Record: found
- Abstract: found
- Article: found