- Record: found
- Abstract: found
- Article: found
CD155/PVR plays a key role in cell motility during tumor cell invasion and migration
Read this article at
Abstract
Background
Invasion is an important early step of cancer metastasis that is not well understood. Developing therapeutics to limit metastasis requires the identification and validation of candidate proteins necessary for invasion and migration.
Methods
We developed a functional proteomic screen to identify mediators of tumor cell invasion. This screen couples Fluorophore Assisted Light Inactivation (FALI) to a scFv antibody library to systematically inactivate surface proteins expressed by human fibrosarcoma cells followed by a high-throughput assessment of transwell invasion.
Results
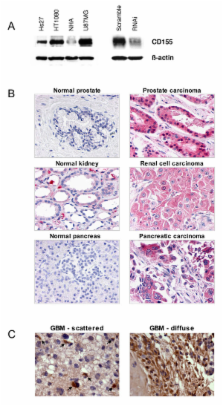
Using this screen, we have identified CD155 (the poliovirus receptor) as a mediator of tumor cell invasion through its role in migration. Knockdown of CD155 by FALI or by RNAi resulted in a significant decrease in transwell migration of HT1080 fibrosarcoma cells towards a serum chemoattractant. CD155 was found to be highly expressed in multiple cancer cell lines and primary tumors including glioblastoma (GBM). Knockdown of CD155 also decreased migration of U87MG GBM cells. CD155 is recruited to the leading edge of migrating cells where it colocalizes with actin and αv-integrin, known mediators of motility and adhesion. Knockdown of CD155 also altered cellular morphology, resulting in cells that were larger and more elongated than controls when plated on a Matrigel substrate.
Related collections
Most cited references34
- Record: found
- Abstract: found
- Article: not found
Molecular classification of cutaneous malignant melanoma by gene expression profiling.
- Record: found
- Abstract: found
- Article: not found
A large-scale RNAi screen in human cells identifies new components of the p53 pathway.
- Record: found
- Abstract: found
- Article: not found