- Record: found
- Abstract: found
- Article: found
Comparative studies of four cumin landraces grown in Egypt
Read this article at
Abstract
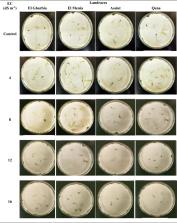
One of the significant aromatic plants applied in food and pharma is cumin. Despite its massive trading in Egypt, there are no comprehensive reports on cumin landraces profile screening. This study aimed to investigate the variation in seeds’ physical and biochemical profiles and genetic diversity as well as assess the efficiency of seeds’ germination under salinity stress. Consequently, during the 2020/2021 growing season, four common cumin seed landraces were gathered from various agro-climatic regions: El Gharbia, El Menia, Assiut, and Qena. Results showed a significant variation in physical profile among the four seeds of landraces. In addition, Assiut had the highest percentage of essential oil at 8.04%, whilst Qena had the largest amount of cumin aldehyde, the primary essential oil component, at 25.19%. Lauric acid was found to be the predominant fatty acid (54.78 to 62.73%). According to ISSR amplification, El Menia presented a negative unique band, whereas other landraces offered a positive band. Additionally, the cumin genotypes were separated into two clusters by the dendrogram, with El Gharbia being located in an entirely separate cluster. There were two sub-clusters within the other cluster: El Menia in one and Assiut and Qena in the other. Moreover, the germination sensitivity to the diverse salinity concentrations (control, 4, 8, 12, and 16 dS/m) findings showed that landraces exhibited varying responses to increased salinity when El Gharbia and El Menia showed a moderate response at four dS/m. Whilst, Qena landraces showed supreme values among other landraces under 12 and 16 dS/m. The majority of the examined features had strong positive associations over a range of salinity levels, according to phenotypic correlation coefficient analysis. To accomplish the aims of sustainable agriculture in Egypt, it would be imperative that the potential breeding program for cumin landraces consider this screening study.
Related collections
Most cited references99
- Record: found
- Abstract: found
- Article: not found
MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods.
- Record: found
- Abstract: found
- Article: not found
Mechanisms of salinity tolerance.

- Record: found
- Abstract: found
- Article: found