- Record: found
- Abstract: found
- Article: found
Evolutionary History of Chemosensory-Related Gene Families across the Arthropoda
Read this article at
Abstract
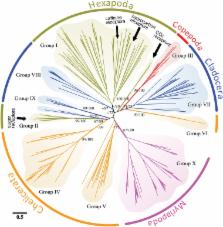
Chemosensory-related gene (CRG) families have been studied extensively in insects, but their evolutionary history across the Arthropoda had remained relatively unexplored. Here, we address current hypotheses and prior conclusions on CRG family evolution using a more comprehensive data set. In particular, odorant receptors were hypothesized to have proliferated during terrestrial colonization by insects (hexapods), but their association with other pancrustacean clades and with independent terrestrial colonizations in other arthropod subphyla have been unclear. We also examine hypotheses on which arthropod CRG family is most ancient. Thus, we reconstructed phylogenies of CRGs, including those from new arthropod genomes and transcriptomes, and mapped CRG gains and losses across arthropod lineages. Our analysis was strengthened by including crustaceans, especially copepods, which reside outside the hexapod/branchiopod clade within the subphylum Pancrustacea. We generated the first high-resolution genome sequence of the copepod Eurytemora affinis and annotated its CRGs. We found odorant receptors and odorant binding proteins present only in hexapods (insects) and absent from all other arthropod lineages, indicating that they are not universal adaptations to land. Gustatory receptors likely represent the oldest chemosensory receptors among CRGs, dating back to the Placozoa. We also clarified and confirmed the evolutionary history of antennal ionotropic receptors across the Arthropoda. All antennal ionotropic receptors in E. affinis were expressed more highly in males than in females, suggestive of an association with male mate-recognition behavior. This study is the most comprehensive comparative analysis to date of CRG family evolution across the largest and most speciose metazoan phylum Arthropoda.
Related collections
Most cited references103
- Record: found
- Abstract: found
- Article: not found
A codon-based model of nucleotide substitution for protein-coding DNA sequences.
- Record: found
- Abstract: found
- Article: not found
The genome of the model beetle and pest Tribolium castaneum.
- Record: found
- Abstract: found
- Article: not found