- Record: found
- Abstract: found
- Article: found
Genome Editing in Plants: Exploration of Technological Advancements and Challenges
Read this article at
Abstract
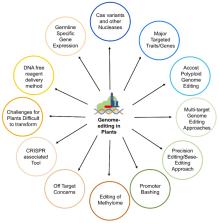
Genome-editing, a recent technological advancement in the field of life sciences, is one of the great examples of techniques used to explore the understanding of the biological phenomenon. Besides having different site-directed nucleases for genome editing over a decade ago, the CRISPR/Cas (clustered regularly interspaced short palindromic repeats/CRISPR-associated protein) based genome editing approach has become a choice of technique due to its simplicity, ease of access, cost, and flexibility. In the present review, several CRISPR/Cas based approaches have been discussed, considering recent advances and challenges to implicate those in the crop improvement programs. Successful examples where CRISPR/Cas approach has been used to improve the biotic and abiotic stress tolerance, and traits related to yield and plant architecture have been discussed. The review highlights the challenges to implement the genome editing in polyploid crop plants like wheat, canola, and sugarcane. Challenges for plants difficult to transform and germline-specific gene expression have been discussed. We have also discussed the notable progress with multi-target editing approaches based on polycistronic tRNA processing, Csy4 endoribonuclease, intron processing, and Drosha ribonuclease. Potential to edit multiple targets simultaneously makes it possible to take up more challenging tasks required to engineer desired crop plants. Similarly, advances like precision gene editing, promoter bashing, and methylome-editing will also be discussed. The present review also provides a catalog of available computational tools and servers facilitating designing of guide-RNA targets, construct designs, and data analysis. The information provided here will be useful for the efficient exploration of technological advances in genome editing field for the crop improvement programs.
Related collections
Most cited references135
- Record: found
- Abstract: found
- Article: not found
Efficient In Vivo Genome Editing Using RNA-Guided Nucleases
- Record: found
- Abstract: found
- Article: not found
Intervening sequences of regularly spaced prokaryotic repeats derive from foreign genetic elements.
- Record: found
- Abstract: found
- Article: not found