- Record: found
- Abstract: found
- Article: found
A Myc-driven self-reinforcing regulatory network maintains mouse embryonic stem cell identity

Read this article at
Abstract
Stem cell identity depends on the integration of extrinsic and intrinsic signals, which directly influence the maintenance of their epigenetic state. Although Myc transcription factors play a major role in stem cell self-renewal and pluripotency, their integration with signalling pathways and epigenetic regulators remains poorly defined. We addressed this point by profiling the gene expression and epigenetic pattern in ESCs whose growth depends on conditional Myc activity. Here we show that Myc potentiates the Wnt/β-catenin signalling pathway, which cooperates with the transcriptional regulatory network in sustaining ESC self-renewal. Myc activation results in the transcriptional repression of Wnt antagonists through the direct recruitment of PRC2 on these targets. The consequent potentiation of the autocrine Wnt/β-catenin signalling induces the transcriptional activation of the endogenous Myc family members, which in turn activates a Myc-driven self-reinforcing circuit. Thus, our data unravel a Myc-dependent self-propagating epigenetic memory in the maintenance of ESC self-renewal capacity.
Abstract
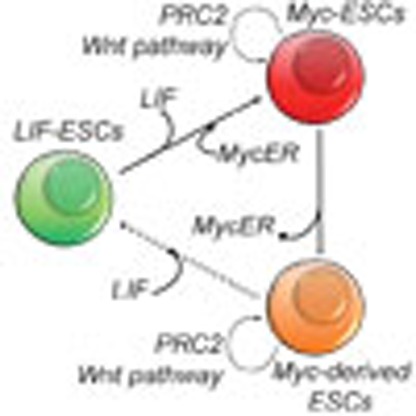 The Myc transcription factor is a major regulator of stem cell (SC) self-renewal and
pluripotency but how this integrates signals from other pathways is unclear. Here,
the authors show that Myc activation triggers epigenetic memory in self renewing embryonic
SCs via PRC2-mediated potentiation of the Wnt pathway.
The Myc transcription factor is a major regulator of stem cell (SC) self-renewal and
pluripotency but how this integrates signals from other pathways is unclear. Here,
the authors show that Myc activation triggers epigenetic memory in self renewing embryonic
SCs via PRC2-mediated potentiation of the Wnt pathway.
Related collections
Most cited references33
- Record: found
- Abstract: found
- Article: not found
Functional expression cloning of Nanog, a pluripotency sustaining factor in embryonic stem cells.
- Record: found
- Abstract: found
- Article: not found
Reconstitution of the mouse germ cell specification pathway in culture by pluripotent stem cells.
- Record: found
- Abstract: found
- Article: not found