- Record: found
- Abstract: found
- Article: found
Structures of membrane proteins
Read this article at
Abstract
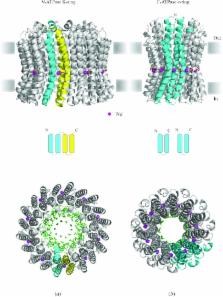
In reviewing the structures of membrane proteins determined up to the end of 2009, we present in words and pictures the most informative examples from each family. We group the structures together according to their function and architecture to provide an overview of the major principles and variations on the most common themes. The first structures, determined 20 years ago, were those of naturally abundant proteins with limited conformational variability, and each membrane protein structure determined was a major landmark. With the advent of complete genome sequences and efficient expression systems, there has been an explosion in the rate of membrane protein structure determination, with many classes represented. New structures are published every month and more than 150 unique membrane protein structures have been determined. This review analyses the reasons for this success, discusses the challenges that still lie ahead, and presents a concise summary of the key achievements with illustrated examples selected from each class.
Related collections
Most cited references325
- Record: found
- Abstract: found
- Article: not found
High-resolution crystal structure of an engineered human beta2-adrenergic G protein-coupled receptor.
- Record: found
- Abstract: found
- Article: not found
Crystal structure of rhodopsin: A G protein-coupled receptor.
- Record: found
- Abstract: found
- Article: not found