- Record: found
- Abstract: found
- Article: found
Genomic Profiling of Childhood Tumor Patient-Derived Xenograft Models to Enable Rational Clinical Trial Design

Read this article at
SUMMARY
Accelerating cures for children with cancer remains an immediate challenge as a result of extensive oncogenic heterogeneity between and within histologies, distinct molecular mechanisms evolving between diagnosis and relapsed disease, and limited therapeutic options. To systematically prioritize and rationally test novel agents in preclinical murine models, researchers within the Pediatric Preclinical Testing Consortium are continuously developing patient-derived xenografts (PDXs)—many of which are refractory to current standard-of-care treatments—from high-risk childhood cancers. Here, we genomically characterize 261 PDX models from 37 unique pediatric cancers; demonstrate faithful recapitulation of histologies and subtypes; and refine our understanding of relapsed disease. In addition, we use expression signatures to classify tumors for TP53 and NF1 pathway inactivation. We anticipate that these data will serve as a resource for pediatric oncology drug development and will guide rational clinical trial design for children with cancer.
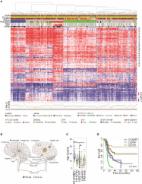
Graphical Abstract

In Brief
Rokita et. al provide an extensively annotated genomic dataset of somatic oncogenic regulation across 37 distinct pediatric malignancies. The 261 patient-derived xenograft models are available to the scientific community, and the genomic annotations will enable rational preclinical agent prioritization and acceleration of therapeutic targets for early-phase pediatric oncology clinical trials.
Related collections
Most cited references43
- Record: found
- Abstract: found
- Article: not found
An algorithm for fast preranked gene set enrichment analysis using cumulative statistic calculation
- Record: found
- Abstract: found
- Article: found
Integrated Molecular Meta-Analysis of 1,000 Pediatric High-Grade and Diffuse Intrinsic Pontine Glioma
- Record: found
- Abstract: found
- Article: not found
Sequencing of neuroblastoma identifies chromothripsis and defects in neuritogenesis genes.
Author and article information
Comments
Comment on this article
 Smart Citations
Smart CitationsSee how this article has been cited at scite.ai
scite shows how a scientific paper has been cited by providing the context of the citation, a classification describing whether it supports, mentions, or contrasts the cited claim, and a label indicating in which section the citation was made.
