- Record: found
- Abstract: found
- Article: found
High-density linkage map construction and identification of loci regulating fruit quality traits in blueberry
Read this article at
Abstract
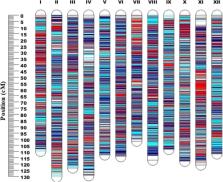
Fruit quality traits play a significant role in consumer preferences and consumption in blueberry ( Vaccinium corymbosum L). The objectives of this study were to construct a high-density linkage map and to identify the underlying genetic basis of fruit quality traits in blueberry. A total of 287 F 1 individuals derived from a cross between two southern highbush blueberry cultivars, ‘Reveille’ and ‘Arlen’, were phenotyped over three years (2016–2018) for fruit quality-related traits, including titratable acidity, pH, total soluble solids, and fruit weight. A high-density linkage map was constructed using 17k single nucleotide polymorphisms markers. The linkage map spanned a total of 1397 cM with an average inter-loci distance of 0.08 cM. The quantitative trait loci interval mapping based on the hidden Markov model identified 18 loci for fruit quality traits, including seven loci for fruit weight, three loci for titratable acidity, five loci for pH, and three loci for total soluble solids. Ten of these loci were detected in more than one year. These loci explained phenotypic variance ranging from 7 to 28% for titratable acidity and total soluble solid, and 8–13% for pH. However, the loci identified for fruit weight did not explain more than 10% of the phenotypic variance. We also reported the association between fruit quality traits and metabolites detected by Proton nuclear magnetic resonance analysis directly responsible for these fruit quality traits. Organic acids, citric acid, and quinic acid were significantly ( P < 0.05) and positively correlated with titratable acidity. Sugar molecules showed a strong and positive correlation with total soluble solids. Overall, the study dissected the genetic basis of fruit quality traits and established an association between these fruit quality traits and metabolites.
Related collections
Most cited references46

- Record: found
- Abstract: found
- Article: found
Fast and accurate short read alignment with Burrows–Wheeler transform

- Record: found
- Abstract: found
- Article: found
The variant call format and VCFtools
- Record: found
- Abstract: not found
- Article: not found