- Record: found
- Abstract: found
- Article: found
Age influences the temporal dynamics of microbiome and antimicrobial resistance genes among fecal bacteria in a cohort of production pigs
Read this article at
Abstract
Background
The pig gastrointestinal tract hosts a diverse microbiome, which can serve to select and maintain a reservoir of antimicrobial resistance genes (ARG). Studies suggest that the types and quantities of antimicrobial resistance (AMR) in fecal bacteria change as the animal host ages, yet the temporal dynamics of AMR within communities of bacteria in pigs during a full production cycle remains largely unstudied.
Results
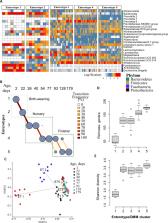
A longitudinal study was performed to evaluate the dynamics of fecal microbiome and AMR in a cohort of pigs during a production cycle; from birth to market age. Our data showed that piglet fecal microbial communities assemble rapidly after birth and become more diverse with age. Individual piglet fecal microbiomes progressed along similar trajectories with age-specific community types/enterotypes and showed a clear shift from E. coli/ Shigella-, Fusobacteria-, Bacteroides-dominant enterotypes to Prevotella-, Megaspheara-, and Lactobacillus-dominated enterotypes with aging. Even when the fecal microbiome was the least diverse, the richness of ARGs, quantities of AMR gene copies, and counts of AMR fecal bacteria were highest in piglets at 2 days of age; subsequently, these declined over time, likely due to age-related competitive changes in the underlying microbiome. ARGs conferring resistance to metals and multi-compound/biocides were detected predominately at the earliest sampled ages.
Related collections
Most cited references88

- Record: found
- Abstract: found
- Article: found
Trimmomatic: a flexible trimmer for Illumina sequence data

- Record: found
- Abstract: found
- Article: found
The Sequence Alignment/Map format and SAMtools

- Record: found
- Abstract: found
- Article: found