- Record: found
- Abstract: found
- Article: found
A genetically modified minipig model for Alzheimer’s disease with SORL1 haploinsufficiency
Read this article at
Summary
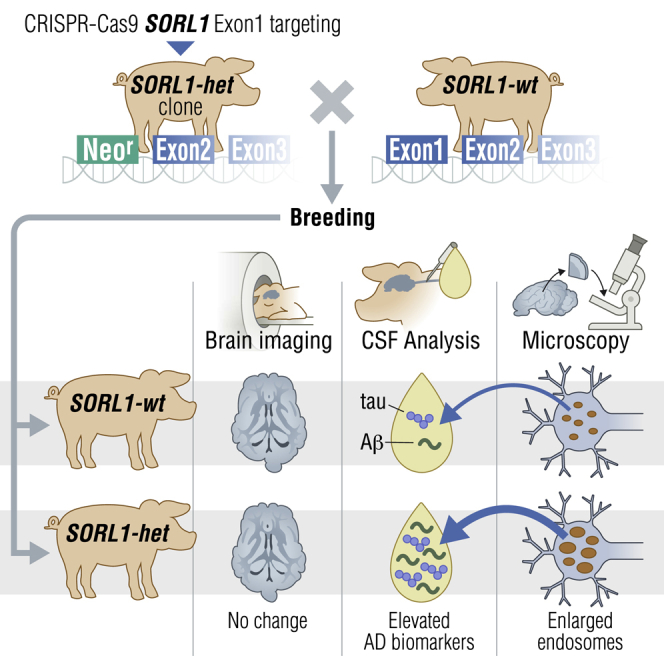
The established causal genes in Alzheimer’s disease (AD), APP, PSEN1, and PSEN2, are functionally characterized using biomarkers, capturing an in vivo profile reflecting the disease’s initial preclinical phase. Mutations in SORL1, encoding the endosome recycling receptor SORLA, are found in 2%–3% of individuals with early-onset AD, and SORL1 haploinsufficiency appears to be causal for AD. To test whether SORL1 can function as an AD causal gene, we use CRISPR-Cas9-based gene editing to develop a model of SORL1 haploinsufficiency in Göttingen minipigs, taking advantage of porcine models for biomarker investigations. SORL1 haploinsufficiency in young adult minipigs is found to phenocopy the preclinical in vivo profile of AD observed with APP, PSEN1, and PSEN2, resulting in elevated levels of β-amyloid (Aβ) and tau preceding amyloid plaque formation and neurodegeneration, as observed in humans. Our study provides functional support for the theory that SORL1 haploinsufficiency leads to endosome cytopathology with biofluid hallmarks of autosomal dominant AD.
Graphical abstract
Highlights
-
•
Minipig model of Alzheimer’s disease by CRISPR knockout of the causal gene SORL1
-
•
Young SORL1 het minipigs phenocopy a preclinical CSF biomarker profile of individuals with AD
-
•
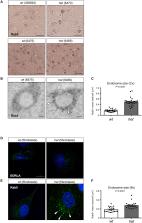
SORL1 haploinsufficiency causes enlarged endosomes similar to neuronal AD pathology
-
•
A minipig model bridging the translational gap between AD mouse models and affected individuals
Abstract
Andersen et al. develop a Göttingen minipig model for Alzheimer’s Disease (AD) with SORL1 haploinsufficiency and demonstrate that their young (<3 years) minipigs have enlarged endosomes and elevated Aβ peptide and tau CSF levels but are unaffected by AD brain pathologies and neurodegeneration, as assessed by PET and MRI scanning methods.
Related collections
Most cited references86
- Record: found
- Abstract: found
- Article: not found