- Record: found
- Abstract: found
- Article: found
PRDM16 binds MED1 and controls chromatin architecture to determine a brown fat transcriptional program
Read this article at
Abstract
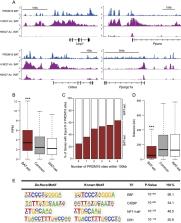
Harms et al. reveal that PRDM16 binding is highly enriched at a broad set of brown fat-selective genes. PRDM16 physically binds to MED1, a component of the Mediator complex, and recruits it to superenhancers at brown fat-selective genes. PRDM16 deficiency in BAT reduces MED1 binding at PRDM16 target sites and causes a fundamental change in chromatin architecture at key brown fat-selective genes.
Abstract
PR (PRD1–BF1–RIZ1 homologous) domain-containing 16 (PRDM16) drives a brown fat differentiation program, but the mechanisms by which PRDM16 activates brown fat-selective genes have been unclear. Through chromatin immunoprecipitation (ChIP) followed by deep sequencing (ChIP-seq) analyses in brown adipose tissue (BAT), we reveal that PRDM16 binding is highly enriched at a broad set of brown fat-selective genes. Importantly, we found that PRDM16 physically binds to MED1, a component of the Mediator complex, and recruits it to superenhancers at brown fat-selective genes. PRDM16 deficiency in BAT reduces MED1 binding at PRDM16 target sites and causes a fundamental change in chromatin architecture at key brown fat-selective genes. Together, these data indicate that PRDM16 controls chromatin architecture and superenhancer activity in BAT.
Related collections
Most cited references36

- Record: found
- Abstract: found
- Article: found
BEDTools: a flexible suite of utilities for comparing genomic features
- Record: found
- Abstract: found
- Article: not found
Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources.
- Record: found
- Abstract: found
- Article: not found