- Record: found
- Abstract: found
- Article: found
FIP200, a ULK-interacting protein, is required for autophagosome formation in mammalian cells
Read this article at
Abstract
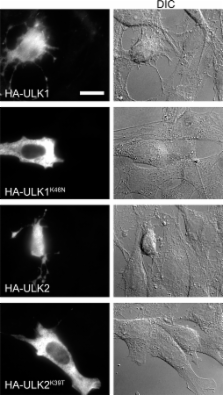
Autophagy is a membrane-mediated intracellular degradation system. The serine/threonine kinase Atg1 plays an essential role in autophagosome formation. However, the role of the mammalian Atg1 homologues UNC-51–like kinase (ULK) 1 and 2 are not yet well understood. We found that murine ULK1 and 2 localized to autophagic isolation membrane under starvation conditions. Kinase-dead alleles of ULK1 and 2 exerted a dominant-negative effect on autophagosome formation, suggesting that ULK kinase activity is important for autophagy. We next screened for ULK binding proteins and identified the focal adhesion kinase family interacting protein of 200 kD (FIP200), which regulates diverse cellular functions such as cell size, proliferation, and migration. We found that FIP200 was redistributed from the cytoplasm to the isolation membrane under starvation conditions. In FIP200-deficient cells, autophagy induction by various treatments was abolished, and both stability and phosphorylation of ULK1 were impaired. These results suggest that FIP200 is a novel mammalian autophagy factor that functions together with ULKs.
Related collections
Most cited references90
- Record: found
- Abstract: found
- Article: not found
RADIOAUTOGRAPHIC STUDIES OF CHOLINE INCORPORATION INTO PERIPHERAL NERVE MYELIN
- Record: found
- Abstract: found
- Article: not found
Systematic identification of protein complexes in Saccharomyces cerevisiae by mass spectrometry.
- Record: found
- Abstract: found
- Article: not found