- Record: found
- Abstract: found
- Article: found
Genome-wide identification and analysis of GRAS transcription factors in the bottle gourd genome
Read this article at
Abstract
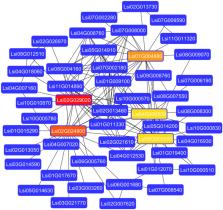
GRAS genes belong to the plant-specific transcription factors (TF’s) family that are known to be involved in plant growth and development. In this study, we have identified 37 genes from the bottle gourd genome that encodes for GRAS TF’s. Except for the SCLA, we were able to identify at least one gene from each of the 17 subfamilies. Gene structure and chromosomal analysis showed that maximum seven genes are present on Chr7 followed by six genes on Chr1. The subcellular location analysis revealed that most of the genes were localized in the nucleus, except for a few in chloroplast and mitochondria. Additionally, we have identified one tandem gene duplication event on Chr7 and three major motifs that were present in all the GRAS genes. Furthermore, the protein–protein interaction prediction and gene expression analysis showed five candidate hub-genes interact with various other genes and thus probably control the expression of interacting partners in different plant tissues. Overall, this study provides a comprehensive analysis of GRAS transcription factors in bottle gourd genome which could be further extended to other vegetable crops.
Related collections
Most cited references31
- Record: found
- Abstract: found
- Article: not found
SMART, a simple modular architecture research tool: identification of signaling domains.
- Record: found
- Abstract: found
- Article: not found
The SHORT-ROOT gene controls radial patterning of the Arabidopsis root through radial signaling.
- Record: found
- Abstract: found
- Article: not found