- Record: found
- Abstract: found
- Article: not found
The Function, Structure, and Origins of the ER Membrane Protein Complex
Read this article at
Abstract
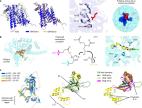
The endoplasmic reticulum (ER) is the site of membrane protein insertion, folding, and assembly in eukaryotes. Over the past few years, a combination of genetic and biochemical studies have implicated an abundant factor termed the ER membrane protein complex (EMC) in several aspects of membrane protein biogenesis. This large nine-protein complex is built around a deeply conserved core formed by the EMC3–EMC6 subcomplex. EMC3 belongs to the universally conserved Oxa1 superfamily of membrane protein transporters, whereas EMC6 is an ancient, widely conserved obligate partner. EMC has an established role in the insertion of transmembrane domains (TMDs) and less understood roles during the later steps of membrane protein folding and assembly. Several recent structures suggest hypotheses about the mechanism(s) of TMD insertion by EMC, with various biochemical and proteomics studies beginning to reveal the range of EMC's membrane protein substrates.
Related collections
Most cited references137
- Record: found
- Abstract: found
- Article: not found
Predicting transmembrane protein topology with a hidden Markov model: application to complete genomes.
- Record: found
- Abstract: found
- Article: not found
Molecular chaperones in protein folding and proteostasis.
- Record: found
- Abstract: found
- Article: found
