- Record: found
- Abstract: found
- Article: found
Detection of Haitian ctxB7 & tcpA alleles in Vibrio cholerae O1 El Tor biotype in Puri, Odisha, India
letter
Read this article at
There is no author summary for this article yet. Authors can add summaries to their articles on ScienceOpen to make them more accessible to a non-specialist audience.
Abstract
Sir,
Vibrio cholerae O1, the causative agent of the majority of cholera outbreaks is classified
into two biotypes, classical and El Tor1 based on the assays such as chicken cell
agglutination (CCA), Voges-Proskauer (VP) reaction, sheep erythrocyte lysis and polymyxin
B susceptibility test2. Over the decades, three variants of V. cholerae O1 El Tor
biotype have been reported globally such as Maltab variants in 2002, Mozambiqe variant
in 2004-2005 and the altered El Tor type (El Tor variant) carrying classical cholera
toxin since 20023
4
5.
The clinical manifestation of cholera is caused by cholera toxin, the major virulence
factor encoded by ctxAB gene located on the CTX prophage integrated on the V. cholerae
chromosome. Classical V. cholerae contains classical type ctxB and El Tor contains
El Tor type ctxB. V. cholerae O1 with a typical El Tor phenotypes (resistant to 50
Units of polymyxin B and positive for CCA and VP test) but carrying ctxB classical
(ctxB1 genotype) is designated as El Tor variant6. On chromosome-1, the single-nucleotide
polymorphism (SNP) at codon 19 results in replacement of the classical cholera toxin
B histidine with asparagine residue resulting in a new ctxB genotype which is named
as ctxB7
7
8.
El Tor variant carrying this new type variant ctxB7 has been reported throughout the
world but these were highlighted after the huge epidemic in Haiti in 2010 and designated
as Haitian variant9. Genomic analysis of Haitian V. cholerae O1 strains revealed mutations
in different segments of chromosome including tcpA (toxin co-regulated pilus) allele10.
Haitian variant gradually spread throughout the world causing cholera outbreaks with
predominance11
12
13. Here, we report the detection of V. cholerae O1 harbouring Haitian variant ctxB
and tcpA associated with polymyxin B (pb, 50 U) susceptibility in Puri district, Odisha,
India, during 2014-2015. This study was a part of the diarrhoea surveillance programme
in Infectious Disease Hospital (IDH) Puri.
During the study, 170 rectal swab samples (72 in 2014 and 98 in 2015) were collected
from hospitalized acute diarrhoea patients. The samples were transported to the ICMR-Regional
Medical Research Center, Bhubaneswar for isolation of V. cholerae. The samples were
incubated for six hours in alkaline peptone water and cultured on thiosulphate-citrate-bile-sucrose
agar (TCBS, BD, USA) followed by biochemical analysis and serotyping14 with polyvalent
O1, O139 and mono-specific Ogawa and Inaba antisera (BD, USA). The study protocol
was approved by the Institutional Ethics Committee. Bacteriological analysis of the
rectal swabs revealed 20 samples positive for V. cholerae O1 Ogawa, El Tor biotype.
All these 20 isolates were found positive for VP test and CCA. Antimicrobial susceptibility
test was carried out by Kirby-Bauer disk diffusion method adhering to Clinical and
Laboratory Standard Institute guidelines15
16. V. cholerae O1 isolates were found sensitive to drugs such as tetracycline, trimethoprim/sulphamethoxazole,
chloramphenicol, neomycin, gentamicin, ciprofloxacin, norfloxacin, ofloxacin, doxycycline
and azithromycin; and resistant to ampicillin, erythromycin, co-trimoxazole, nalidixic
acid and furazolidone. Of the total 20 isolates, 15 (75%) were found sensitive to
Pb (50 units). The genetic analysis of isolated V. cholerae isolates included PCR
assays for detection of wbe, ctxA and tcpA. Haitian variants ctxB and tcpA were amplified
using the primer pairs as described previously10
17. PCR analysis revealed that all the 20 isolates were positive for ctxA, tcpA (El
Tor) and rfb genes. The presence of ctxA (301 bp)in all V. cholerae isolates confirmed
their toxigenic potential, wbe (192 bp) provided the molecular evidence for O1 serogroup
and tcpA El Tor (469 bp) determined the El Tor type. Further biotype-specific CTX
prophage repressor rstR was amplified with El Tor-specific primers18, indicating the
presence of El Tor rstR in all these isolates. All V. cholerae isolates were found
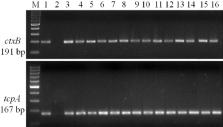
to carry Haitian variant tcpA allele [(167 bp) Figure] which confirmed that tcpA had
a single base substitution at 266-nt position as was found in strains from Haiti19.
Double mismatch-amplification-mutation assay (DMAMA)17 detected Haitian variant ctxB
allele (191 bp) in all 20 V. cholerae O1 isolates during the study (Figure).
Figure
PCR analysis showing Haitian variant ctxB and tcpA allele in V. cholerae O1 isolates.
Lane M, 100 bp size ladder; lane1, Haitian control 2010EL-1786; lane 2, classical
control, O395; from lane 3 through lane 16, V. cholerae O1 isolates.
A surveillance study conducted in three hospitals including IDH during 2004-2006 revealed
the incidence rate of V. cholerae as 17.3 per cent where all V. cholerae O1 were El
Tor biotype20. During 2011-2013, following a study on operational feasibility of an
oral cholera vaccine in Satybadi Block of Puri district21, the post-vaccination diarrhoea
surveillance demonstrated the incidence rate of V. cholerae O1 as 3.4 per cent (unpublished
data) and bio-typing of isolates revealed V. cholerae O1 Hybrid and El Tor variants22.
In the present study, 11.8 per cent V. cholerae isolates were found to be Haitian
variants.
The emergence of Haitian variant, tempted researchers to investigate its incidence
and spread causing cholera outbreaks. Two cholera outbreaks in western and southern
Odisha during 2013 and 2016 respectively presented the ctxB7 genotype (unpublished
data). Our previous study depicted that a large cholera outbreak occurred in western
Odisha during 2014 was due to V. cholerae O1 with ctxB7 and retrospective analysis
of laboratory strains revealed that the origin of Haitian variant was since 1999 in
Odisha23. Earlier studies in Kolkata reported V. cholerae with tcpA of Haitian allele,
ctxB7 allele and Pb susceptibility since 2003, 2006 and 2012, respectively10
17
24. In north India, including Delhi, Haryana and Uttar Pradesh, the combination of
Haitian ctxB (ctxB7), classical ctxB (ctxB1) and tcpA of Haitian allele has been reported
since 200825. A cholera outbreak in Maharashtra during 2012 was due to V. cholerae
O1 carrying Haitian variant ctxB
26. The present study showed the combination of 100 per cent Haitian variants ctxB
and tcpA associated with high (75%) susceptibility to Pb without the detection of
ctxB1 allele. This suggested that Haitian strains might have replaced the El Tor variant
in this region as has been reported for El Tor being replaced by El Tor variant strains
during 2008-2009 in Odisha27. Similar to the Kolkata strains24, the high percentage
(75%) of Pb susceptibility in our isolates was a unique phenotypic change contrast
to the El Tor biotype of earlier decades and Haitian variants in Kerala, south India28.
The cryptic change in the genetic backbone is the current evolutionary outcome in
V. cholerae O1 that influences its virulence, transmission and spread. The presence
of hypervirulent Haitian variant in Puri has significance in connection to the spread
of cholera outbreaks. An effective surveillance acts as a resource to develop early
warning to implement prevention and preparedness for disease outbreak. This report
of Haitian variant in coastal Odisha warrants constant surveillance in other parts
of Odisha and India.
Related collections
Most cited references24
- Record: found
- Abstract: found
- Article: not found
Understanding the Cholera Epidemic, Haiti
Renaud Piarroux, Robert Barrais, Benoit Faucher … (2011)
- Record: found
- Abstract: found
- Article: not found
Cholera due to altered El Tor strains of Vibrio cholerae O1 in Bangladesh.
David A. Sack, Firdausi Qadri, Shah Faruque … (2006)
Author and article information
Comments
Comment on this article
scite_
4
0
0
0
 Smart Citations
Smart Citations4
0
0
0
Citing PublicationsSupportingMentioningContrasting
See how this article has been cited at scite.ai
scite shows how a scientific paper has been cited by providing the context of the citation, a classification describing whether it supports, mentions, or contrasts the cited claim, and a label indicating in which section the citation was made.