- Record: found
- Abstract: found
- Article: found
A High Precision Survey of the Molecular Dynamics of Mammalian Clathrin-Mediated Endocytosis
Read this article at
Abstract
The molecular dynamics of clathrin-mediated endocytosis in living cells has been mapped with an approximately ten-fold improvement in temporal accuracy, yielding new insights into the molecular mechanism.
Abstract
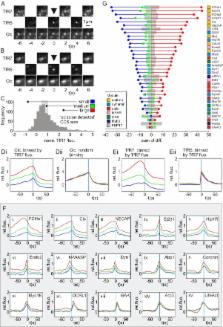
Dual colour total internal reflection fluorescence microscopy is a powerful tool for decoding the molecular dynamics of clathrin-mediated endocytosis (CME). Typically, the recruitment of a fluorescent protein–tagged endocytic protein was referenced to the disappearance of spot-like clathrin-coated structure (CCS), but the precision of spot-like CCS disappearance as a marker for canonical CME remained unknown. Here we have used an imaging assay based on total internal reflection fluorescence microscopy to detect scission events with a resolution of ∼2 s. We found that scission events engulfed comparable amounts of transferrin receptor cargo at CCSs of different sizes and CCS did not always disappear following scission. We measured the recruitment dynamics of 34 types of endocytic protein to scission events: Abp1, ACK1, amphiphysin1, APPL1, Arp3, BIN1, CALM, CIP4, clathrin light chain (Clc), cofilin, coronin1B, cortactin, dynamin1/2, endophilin2, Eps15, Eps8, epsin2, FBP17, FCHo1/2, GAK, Hip1R, lifeAct, mu2 subunit of the AP2 complex, myosin1E, myosin6, NECAP, N-WASP, OCRL1, Rab5, SNX9, synaptojanin2β1, and syndapin2. For each protein we aligned ∼1,000 recruitment profiles to their respective scission events and constructed characteristic “recruitment signatures” that were grouped, as for yeast, to reveal the modular organization of mammalian CME. A detailed analysis revealed the unanticipated recruitment dynamics of SNX9, FBP17, and CIP4 and showed that the same set of proteins was recruited, in the same order, to scission events at CCSs of different sizes and lifetimes. Collectively these data reveal the fine-grained temporal structure of CME and suggest a simplified canonical model of mammalian CME in which the same core mechanism of CME, involving actin, operates at CCSs of diverse sizes and lifetimes.
Author Summary
The molecular machinery of clathrin-mediated endocytosis concentrates receptors at the cell surface in a patch of membrane that curves into a vesicle, pinches off, and internalizes membrane cargo and a tiny volume of extracellular fluid. We know that dozens of proteins are involved in this process, but precisely when and where they act remains poorly understood. Here we used a fluorescence imaging assay to detect the moment of scission in living cells and used this as a reference point from which to measure the characteristic recruitment signatures of 34 fluorescently tagged endocytic proteins. Pair-wise comparison of these recruitment signatures allowed us to identify seven modules of proteins that were recruited with similar kinetics. For the most part the recruitment signatures were consistent with what was previously known about the proteins' structure and their binding affinities; however, the recruitment signatures for some components (such as some BAR and F-BAR domain proteins) could not have been predicted from existing structural or biochemical data. This study provides a paradigm for mapping molecular dynamics in living cells and provides new insights into the mechanism of clathrin-mediated endocytosis.
Related collections
Most cited references66
- Record: found
- Abstract: found
- Article: not found
BAR domains as sensors of membrane curvature: the amphiphysin BAR structure.
- Record: found
- Abstract: found
- Article: not found
Improving the photostability of bright monomeric orange and red fluorescent proteins.
- Record: found
- Abstract: found
- Article: not found