- Record: found
- Abstract: found
- Article: found
Comparison of critical biomarkers in 2 erectile dysfunction models based on GEO and NOS-cGMP-PDE5 pathway
Read this article at
Abstract
Background:
Erectile dysfunction is a disease commonly caused by diabetes mellitus (DMED) and cavernous nerve injury (CNIED). Bioinformatics analyses including differentially expressed genes (DEGs), enriched functions and pathways (EFPs), and protein-protein interaction (PPI) networks were carried out in DMED and CNIED rats in this study. The critical biomarkers that may intervene in nitric oxide synthase (NOS, predominantly nNOS, ancillary eNOS, and iNOS)-cyclic guanosine monophosphate (cGMP)-phosphodiesterase 5 enzyme (PDE5) pathway, an important mechanism in erectile dysfunction treatment, were then explored for potential clinical applications.
Methods:
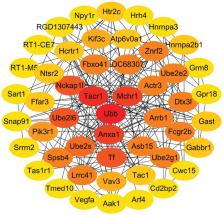
GSE2457 and GSE31247 were downloaded. Their DEGs with a |logFC (fold change)| > 0 were screened out. Database for Annotation, Visualization and Integrated Discovery (DAVID) online database was used to analyze the EFPs in Gene Ontology enrichment and Kyoto Encyclopedia of Genes and Genomes networks based on down-regulated and up-regulated DEGs respectively. PPI analysis of 2 datasets was performed in Search Tool for the Retrieval of Interacting Genes/Proteins (STRING) and Cytoscape. Interactions with an average score greater than 0.9 were chosen as the cutoff for statistical significance.
Results:
From a total of 1710 DEGs in GSE2457, 772 were down-regulated and 938 were up-regulated, in contrast to the 836 DEGs in GSE31247, from which 508 were down-regulated and 328 were up-regulated. The 25 common EFPs such as aging and response to hormone were identified in both models. PPI results showed that the first 10 hub genes in DMED were all different from those in CNIED.
Related collections
Most cited references70
- Record: found
- Abstract: found
- Article: not found
Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources.

- Record: found
- Abstract: found
- Article: found
NCBI GEO: archive for functional genomics data sets—update

- Record: found
- Abstract: found
- Article: found