- Record: found
- Abstract: found
- Article: found
Analysis of Gut Microbiota Signature and Microbe-Disease Progression Associations in Locally Advanced Non-Small Cell Lung Cancer Patients Treated With Concurrent Chemoradiotherapy
Read this article at
Abstract
Purpose
To evaluate the association of gut microbiome signature and disease progression in locally advanced non-small cell lung cancer (LA-NSCLC) patients treated with concurrent chemoradiotherapy (CCRT) by fecal metagenome analysis.
Methods
Metagenome-wide association studies on baseline fecal samples from 18 LA-NSCLC patients before CCRT and 13 controls from healthy first-degree relatives were performed. Among the 18 LA-NSCLC patients, six patients were defined as the long progression-free survival (long-PFS) group (PFS≥11 months) while another 12 were in the short-PFS group (PFS<11 months). Alpha diversity, taxonomic composition, and Kyoto Encyclopedia of Genes and Genomes (KEGG) functional pathways were compared between groups.
Results
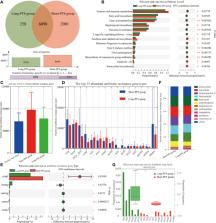
The Firmicutes/Bacteroidetes value of long-PFS group was higher than those of short-PFS (p=0.073) and healthy individual groups (p=0.009). Meanwhile, long-PFS group had significantly higher diversities in Fungi, Archaea, and Viruses than short-PFS group. The KEGG pathways overrepresented in short-PFS group included fructose and mannose metabolism (p=0.028), streptomycin biosynthesis (p=0.028), acarbose and validamycin biosynthesis (p=0.013), ribosome biogenesis in eukaryotes (p=0.035), biosynthesis of vancomycin group antibiotics (p=0.004), apoptosis-fly (p=0.044), and tetracycline biosynthesis (p=0.044), while those overrepresented in long-PFS group included fatty acid biosynthesis (p=0.035), fatty acid metabolism (p=0.008), vancomycin resistance (p=0.008), longevity regulating pathway-worm (p=0.028), type II diabetes mellitus (p=0.004), and viral carcinogenesis (p=0.003). Further analysis of antibiotic resistome demonstrated that the short-PFS group had a trend with more antibiotic resistance genes than healthy control (p=0.070) and long-PFS groups (p=0.218). The vancomycin resistance sequences were significantly enriched in the long-PFS group compared to the short-PFS group (p=0.006).
Conclusions
The baseline gut microbiome composition and functionality might be associated with PFS in LA-NSCLC treated with CCRT. The outcome of CCRT might be modulated through bacterial metabolic pathways. The antibiotic resistance genes might play a role in disease progression and provide potential information on the relationship between the use of antibiotics and treatment efficacy of CCRT in LA-NSCLC.
Related collections
Most cited references53

- Record: found
- Abstract: found
- Article: found
fastp: an ultra-fast all-in-one FASTQ preprocessor
- Record: found
- Abstract: found
- Article: not found
Fast and sensitive protein alignment using DIAMOND.

- Record: found
- Abstract: found
- Article: found